de_genes <- Seurat::FindAllMarkers(sample, only.pos = TRUE)
p <- SCpubr::do_GroupwiseDEHeatmap(sample = sample,
de_genes = de_genes)
p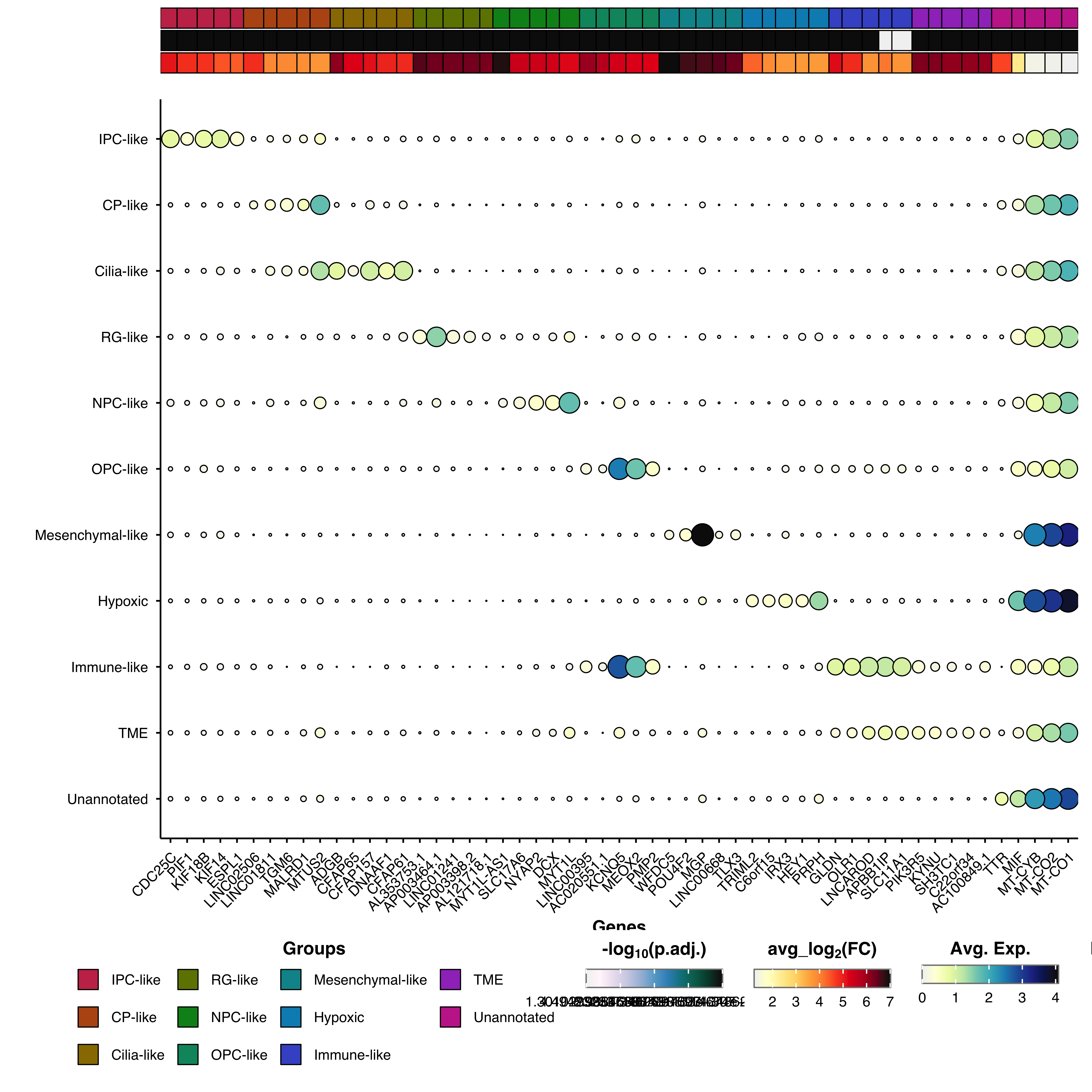
Groupwise DE heatmaps visualize the results of Seurat::FindAllMarkers() as a multi-panel dot plot showing adjusted p-values, fold changes, and mean expression per cluster.
First run FindAllMarkers():
de_genes <- Seurat::FindAllMarkers(sample, only.pos = TRUE)
p <- SCpubr::do_GroupwiseDEHeatmap(sample = sample,
de_genes = de_genes)
p
p <- SCpubr::do_GroupwiseDEHeatmap(sample = sample,
de_genes = de_genes,
top_genes = 3)
p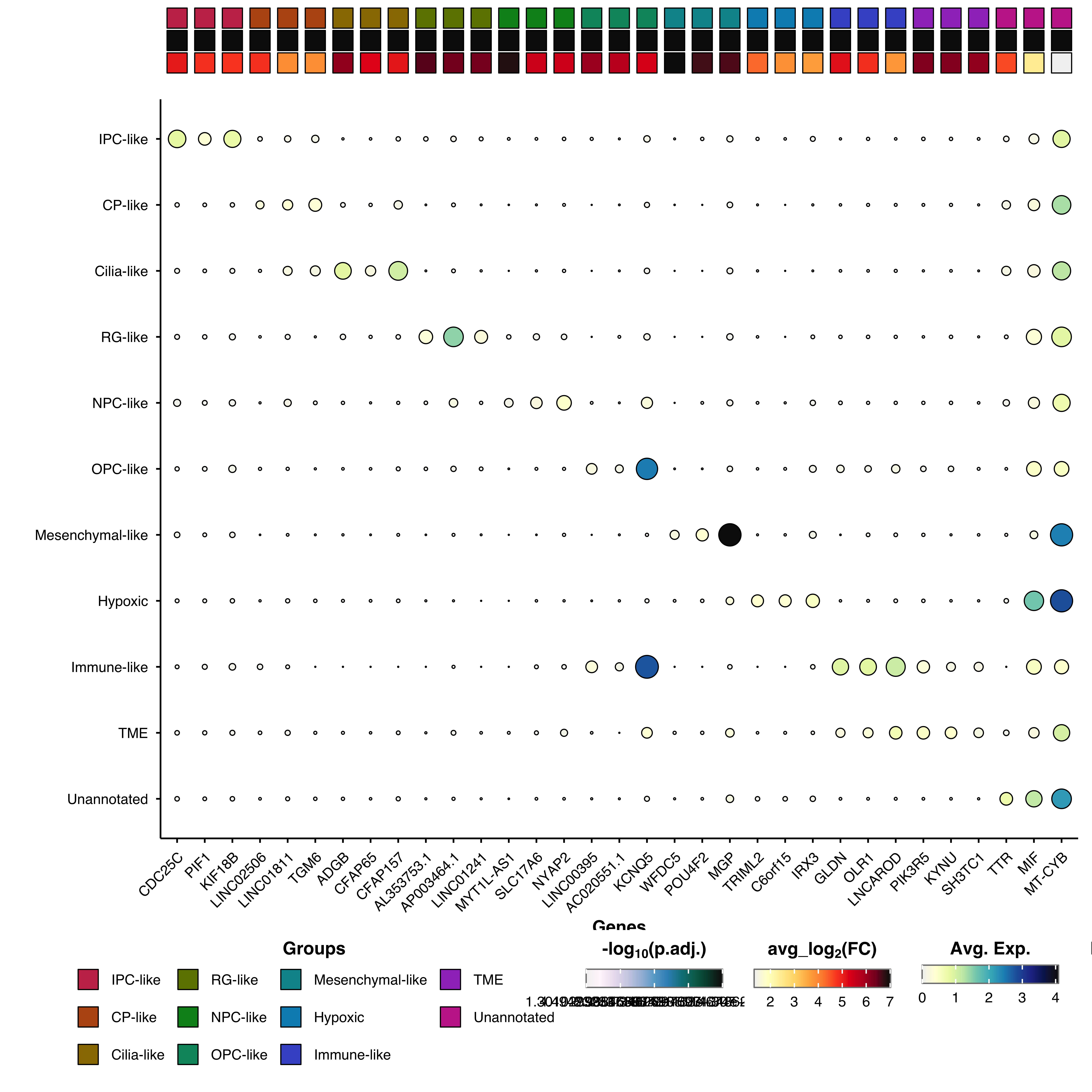
p <- SCpubr::do_GroupwiseDEHeatmap(sample = sample,
de_genes = de_genes,
p.cutoff = 0.01)
p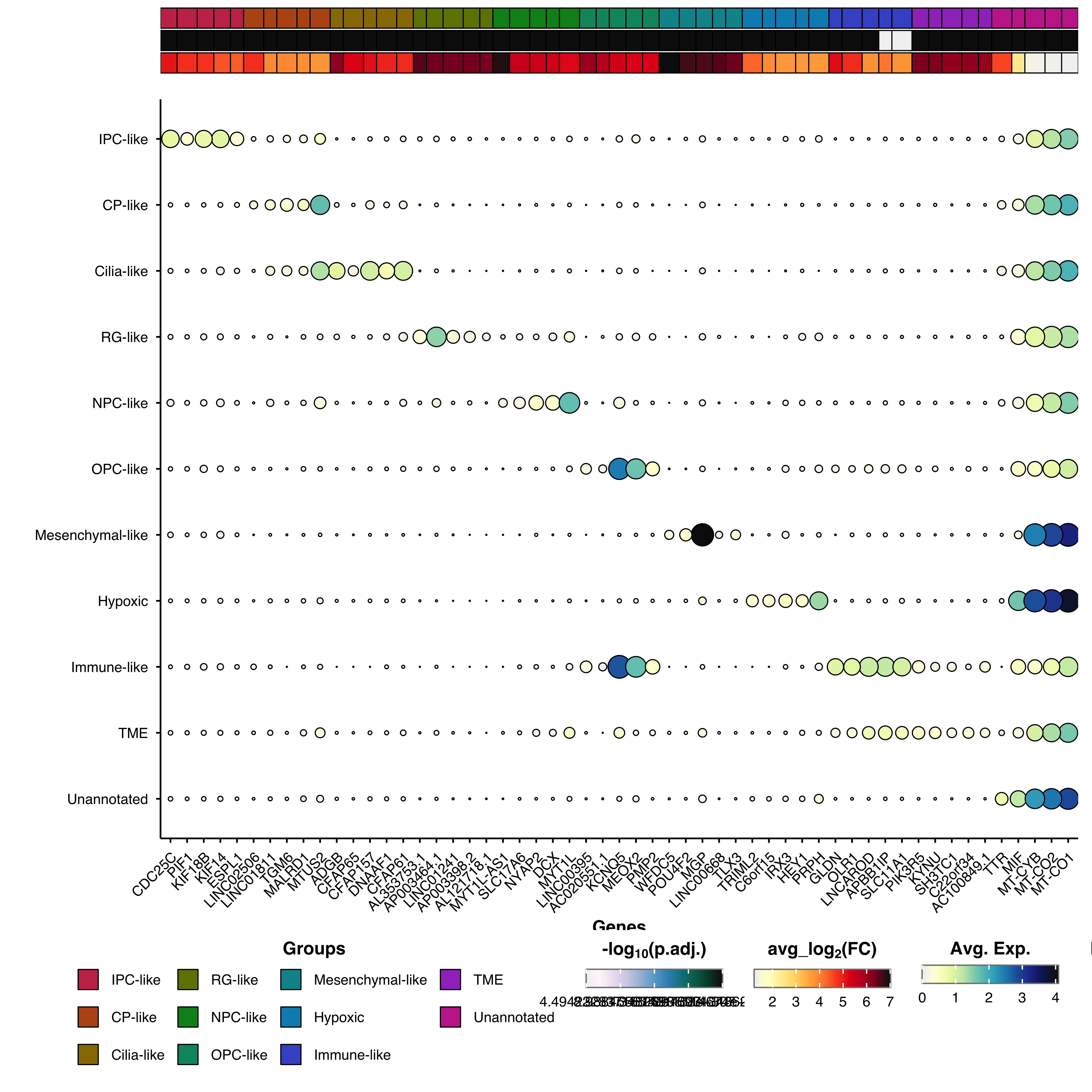
p <- SCpubr::do_GroupwiseDEHeatmap(sample = sample,
de_genes = de_genes,
dot.scale = 6)
p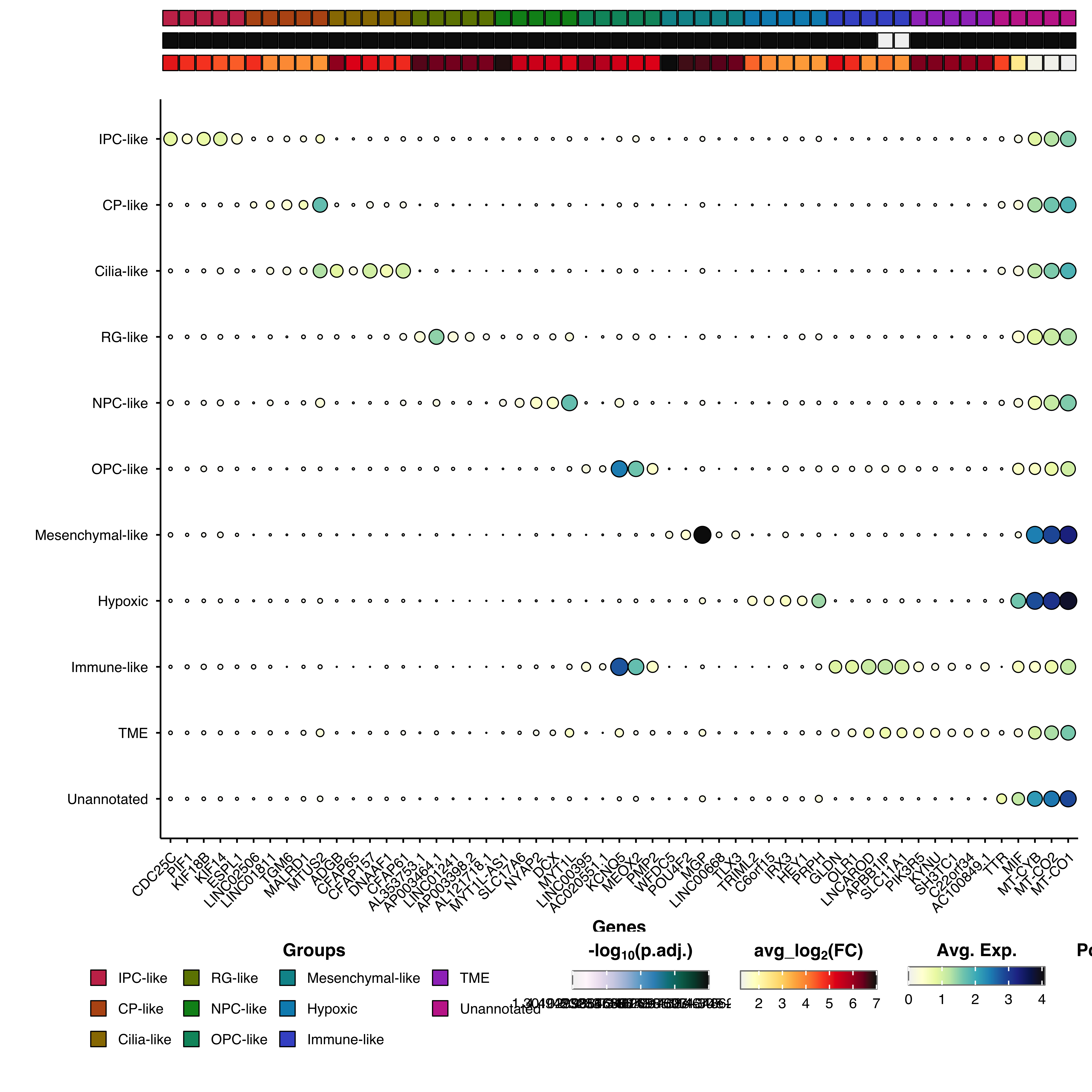
For parameters shared across many functions (color palettes, typography, legend styling), see Shared features.
| Parameter | Description | Default |
|---|---|---|
de_genes |
Output from FindAllMarkers()
|
— |
top_genes |
Top DE genes per group | 5 |
p.cutoff |
Adjusted p-value threshold | 0.05 |
dot.scale |
Dot size multiplier | 8 |