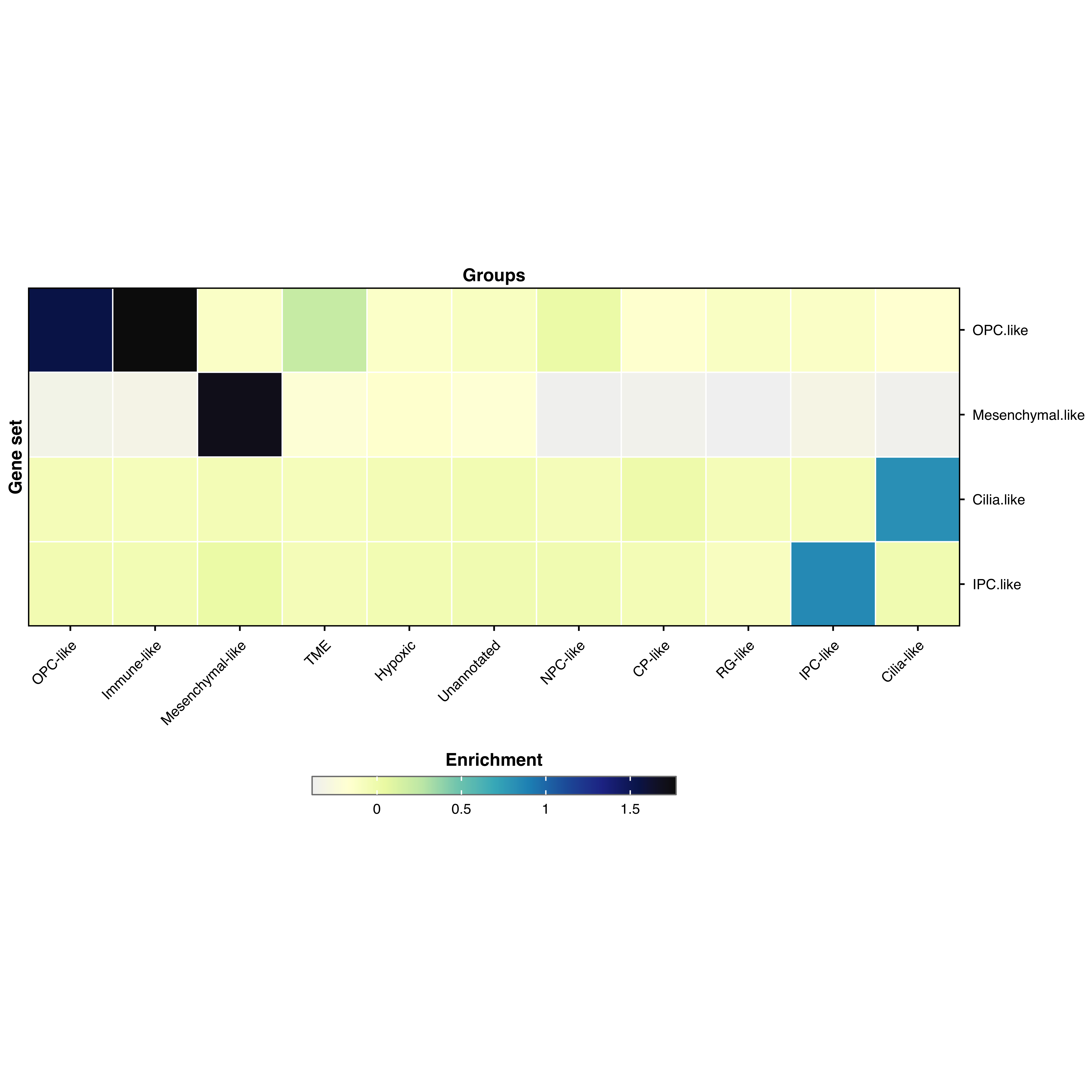
Enrichment heatmaps display averaged enrichment scores for gene sets across groups. This function computes enrichment scores using Seurat’s AddModuleScore (or UCell) and aggregates them by metadata variables.
Basic usage
gene_lists <- list ( "IPC.like" = c ( "CDC25C" , "KIF18B" , "KIF14" , "CENPE" ) ,
"Cilia.like" = c ( "DNAAF1" , "ADGB" , "CFAP61" , "CFAP157" ) ,
"OPC.like" = c ( "KCNQ5" , "MEOX2" , "MBP" , "SNTG1" ) ,
"Mesenchymal.like" = c ( "S100A1" , "MGP" , "TNNT1" , "H2AFJ" )
) p <- SCpubr :: do_EnrichmentHeatmap ( sample = sample , input_gene_list = gene_lists ,
flip = TRUE )
p
Multiple grouping variables
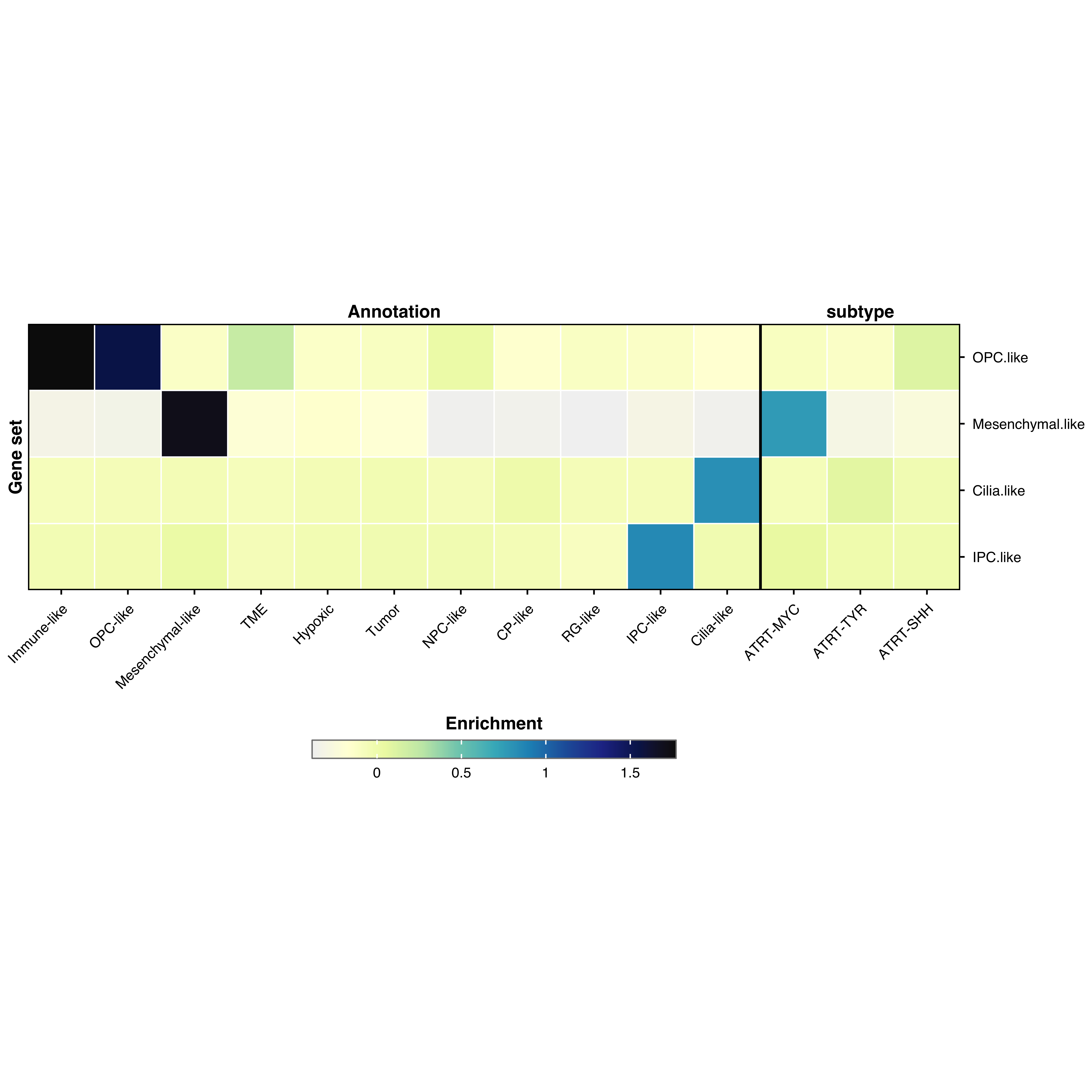
p <- SCpubr :: do_EnrichmentHeatmap ( sample = sample , input_gene_list = gene_lists ,
group.by = c ( "Annotation" , "subtype" ) ,
flip = TRUE )
p
Scoring methods
Use flavor = "Seurat" or flavor = "UCell" to toggle different scoring methods.
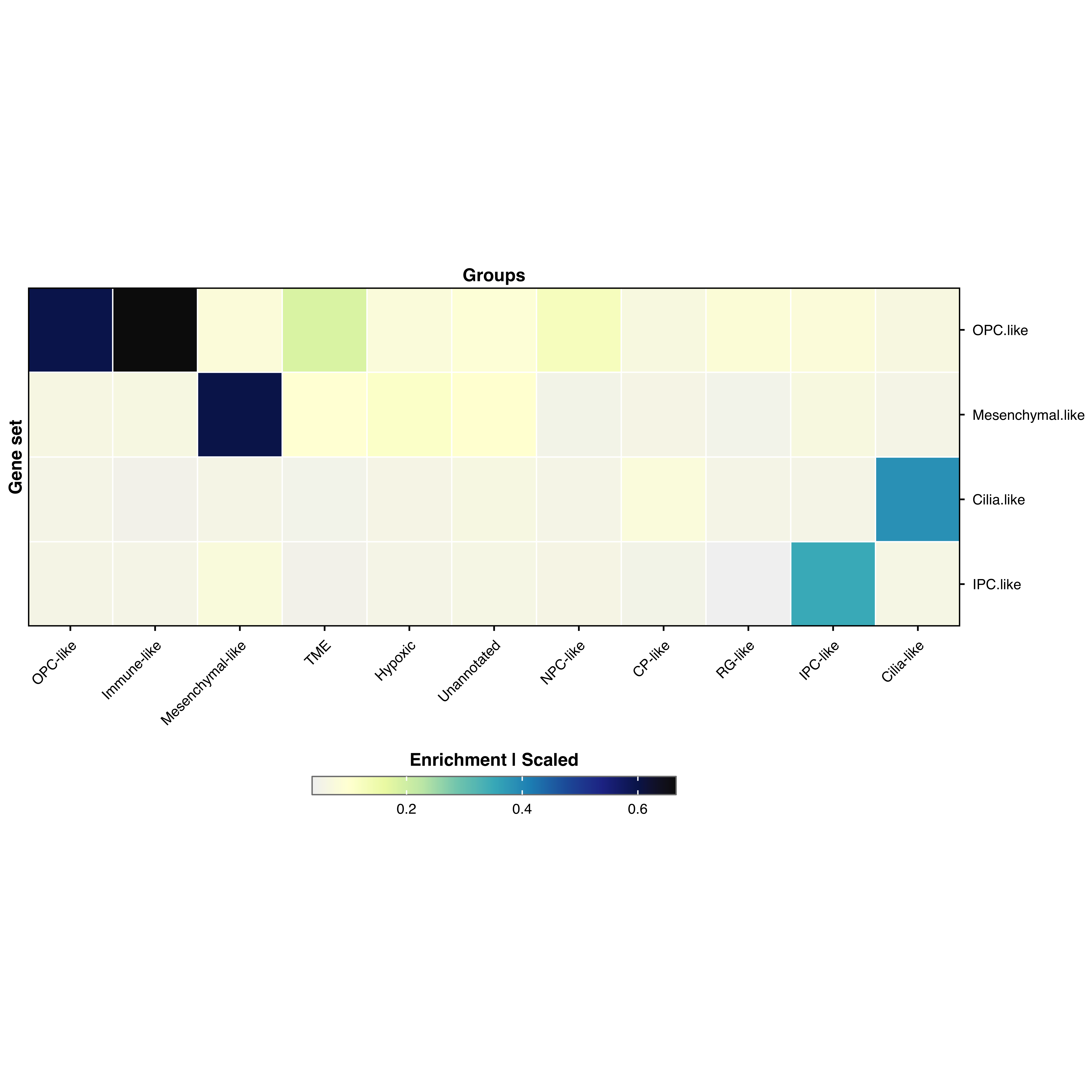
Scale scores
Transform scores to 0-1 range for easier comparison:
p <- SCpubr :: do_EnrichmentHeatmap ( sample = sample , input_gene_list = gene_lists ,
scale_scores = TRUE ,
flip = TRUE )
p
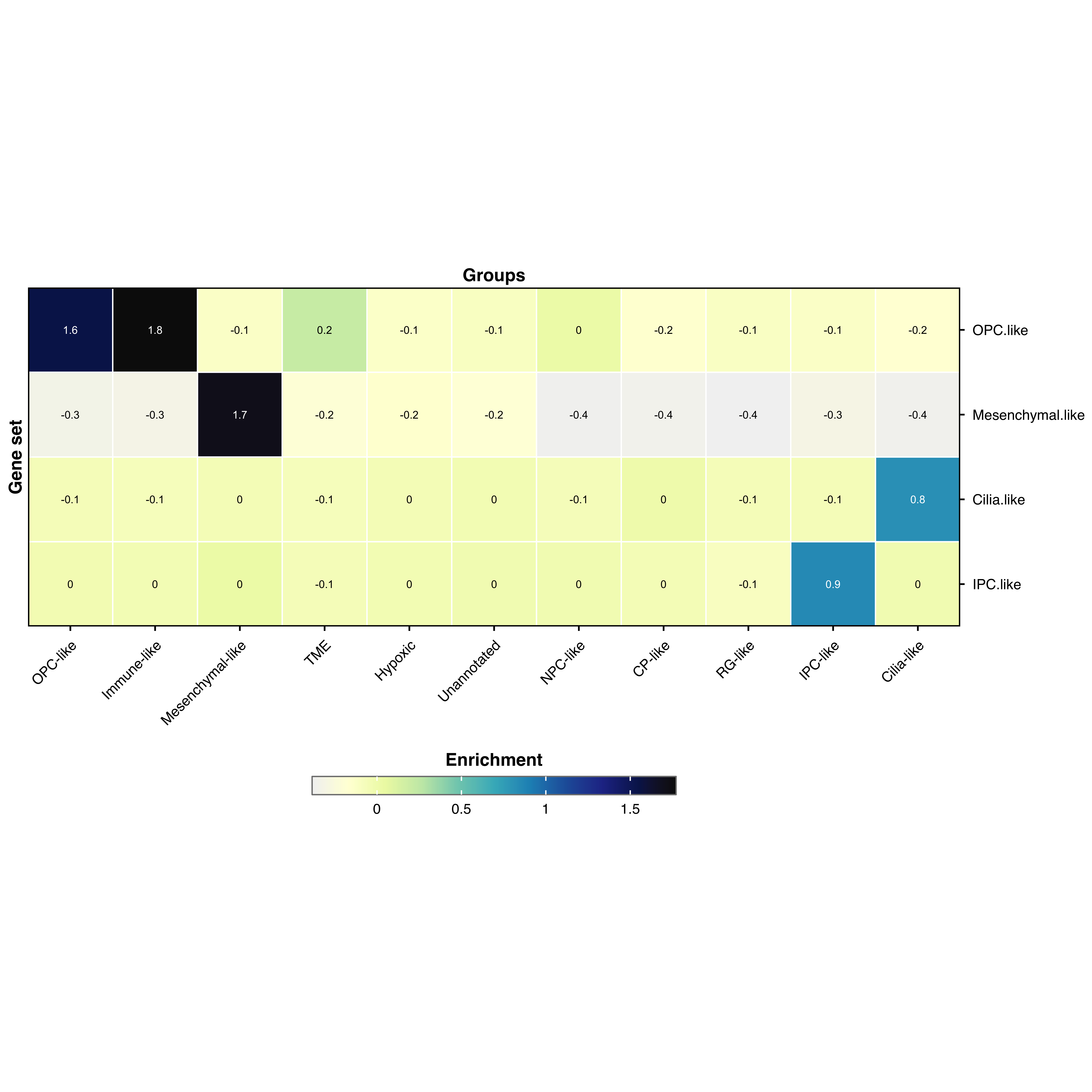
Display values
p <- SCpubr :: do_EnrichmentHeatmap ( sample = sample , input_gene_list = gene_lists ,
values.show = TRUE ,
values.threshold = 0.5 ,
flip = TRUE )
p
Return enriched object
Get the Seurat object with computed enrichment scores:
result <- SCpubr :: do_EnrichmentHeatmap ( sample = sample , input_gene_list = gene_lists ,
return_object = TRUE )
p <- result $ plot sample_enriched <- result $ object
Parameter reference
For parameters shared across many functions (color palettes, typography, legend styling), see Shared features .
Core parameters
input_gene_listNamed list of gene signatures
—
Scoring method
flavorScoring method ("Seurat" or "UCell")
"Seurat"
nbinNumber of bins (Seurat)
24
ctrlControl genes per bin (Seurat)
100
ncoresCPU cores for UCell
1
storeRanksStore ranks for faster UCell
TRUE
scale_scoresScale scores to 0-1 range
FALSE
Clustering & ordering
clusterHierarchically cluster
TRUE
features.orderCustom gene set order
NULL
groups.orderCustom group order
NULL
Output
return_objectReturn Seurat object with scores
FALSE