Ranked expression heatmaps display gene expression ordered by a dimensional reduction component . Cells are ranked along a PC, UMAP dimension, or trajectory score, revealing expression patterns along continuous gradients.
Basic usage
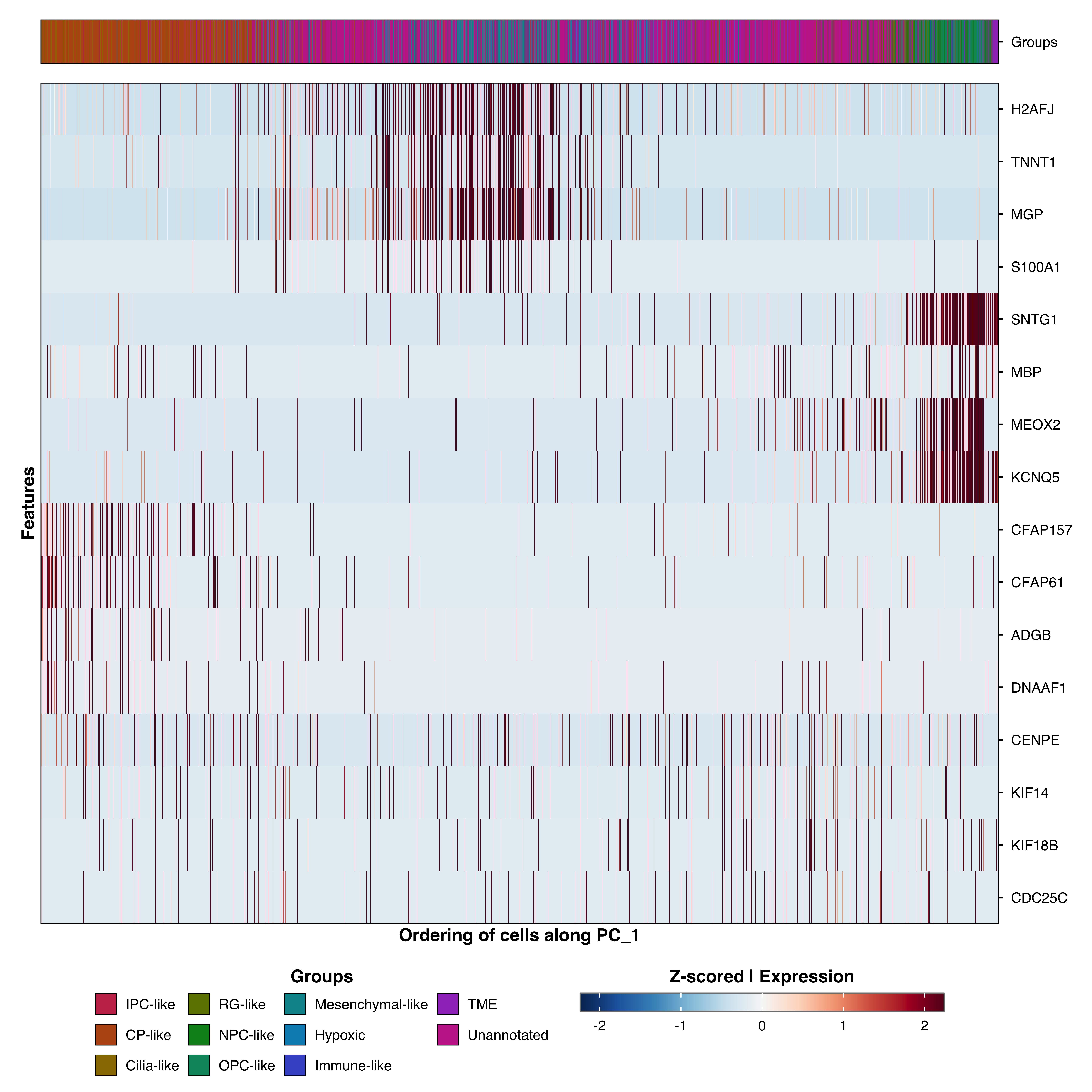
Rank cells by the first PC component:
genes <- c ( "CDC25C" , "KIF18B" , "KIF14" , "CENPE" , "DNAAF1" , "ADGB" , "CFAP61" , "CFAP157" ,
"KCNQ5" , "MEOX2" , "MBP" , "SNTG1" ,
"S100A1" , "MGP" , "TNNT1" , "H2AFJ" )
p <- SCpubr :: do_RankedExpressionHeatmap ( sample = sample , features = genes ,
reduction = "pca" ,
dims = 1 )
p #> $PC_1
Multiple dimensions
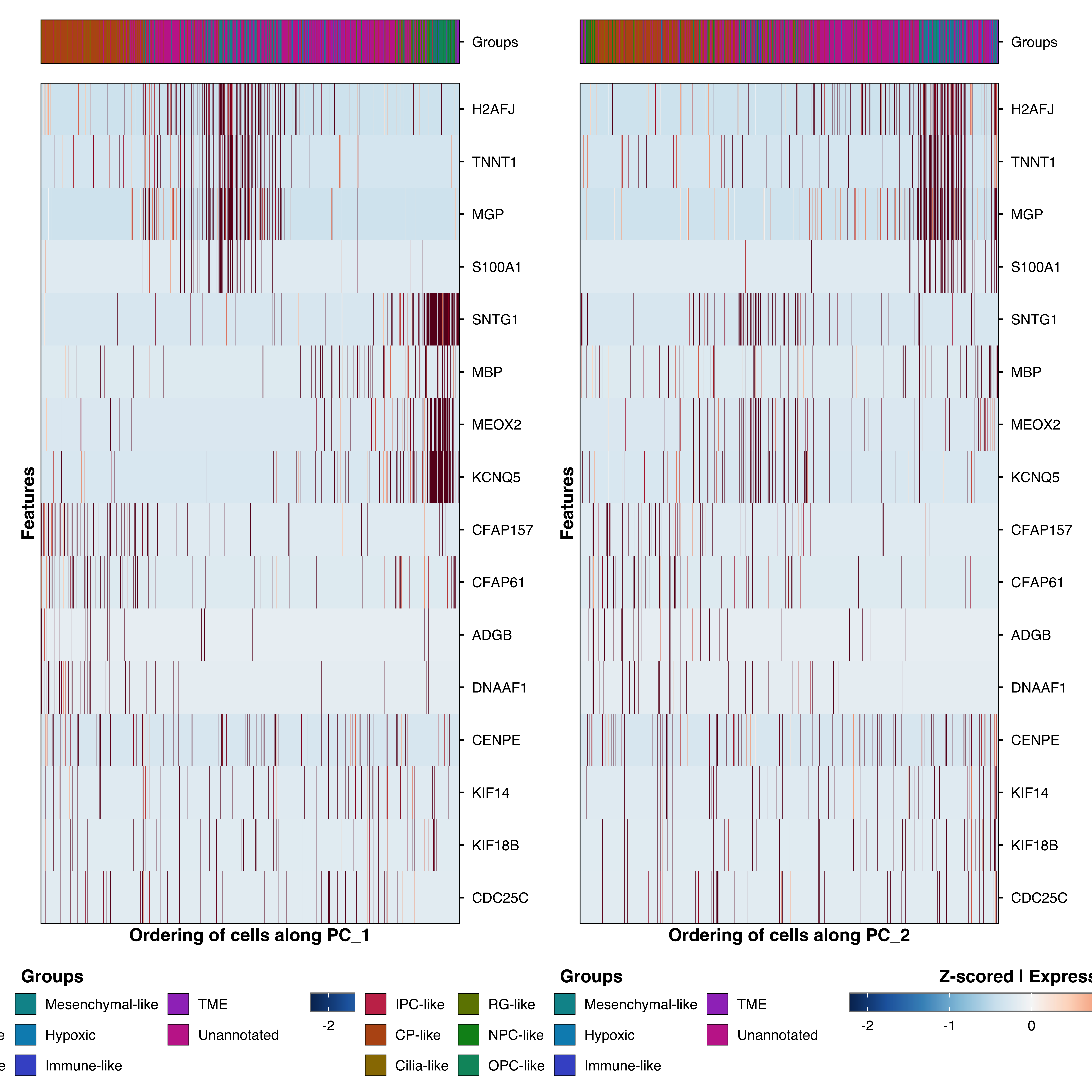
Generate one heatmap per dimension:
p <- SCpubr :: do_RankedExpressionHeatmap ( sample = sample , features = genes ,
reduction = "pca" ,
dims = 1 : 2 )
# Access individual plots p $ PC_1 | p $ PC_2
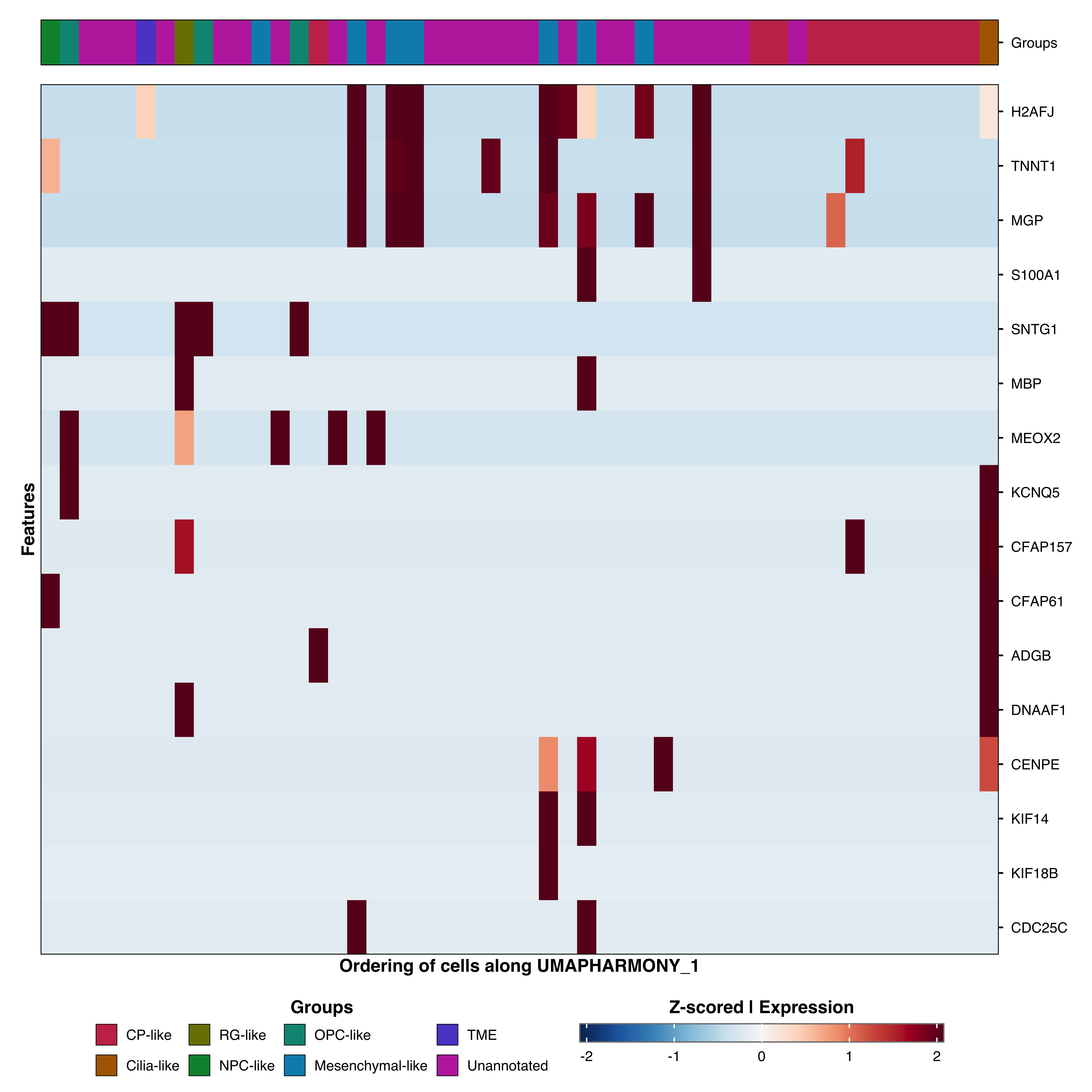
Subsampling
Reduce cells for faster plotting:
Parameter reference
For parameters shared across many functions (color palettes, typography, legend styling), see Shared features .
Core parameters
featuresGenes to display
—
dimsDimensions to rank cells by
1:2
Appearance
main.heatmap.sizeMain heatmap proportion (0-1)
0.95
subsampleMax cells to display
2500
interpolateSmooth heatmap
FALSE